This week’s Spotlight is on Dr. Adam Gazzaley of UC San Francisco, where he is the founding director of the Neuroscience Imaging Center and an Associate Professor in Neurology, Physiology and Psychiatry. His work was featured in Nature in September 2013.
 NVIDIA: Adam, how are you using GPU computing in your research?
NVIDIA: Adam, how are you using GPU computing in your research?
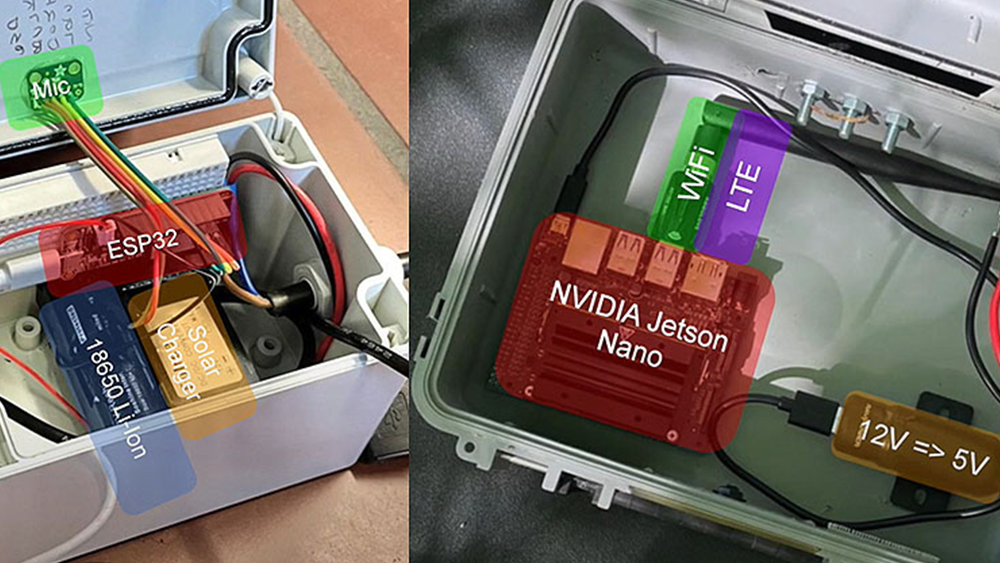
Adam: We are working with a distributed team (UCSF, Stanford, UCSD and Eye Vapor) to CUDA-enable EEG (electroencephalography) processing to increase the fidelity of real-time brain activity recordings.
The goal is to more accurately represent the brain sources and neural networks, as well as to perform real-time artifact correction and mental state decoding. Not only will this improve the visualization capabilities, but more importantly, it will move EEG closer to being a real-time scientific tool.
Where CUDA and the GPU really excel is with very intense computations that use large matrices. We generate that type of data when we’re recording real-time brain activity across many electrodes.

NVIDIA: Describe the hardware/software platform currently in use by the development team.
Adam: We primarily use Python, MATLAB and C/C++. Our software is routinely executed on a range of platforms, including Linux (running Fedora 18), Windows 7, and Mac OS (Snow Leopard and Lion).Hardware we currently make use of includes NVIDIA Tesla K20s (for calculations), NVIDIA Quadro 5000s (for visualization) and two Intel Quad-core CPUs.
We use Microsoft Visual Studio 2010 x64 with CUDA 5.0, with the TCC driver for the Tesla GPUs. The Nvidia Nsight debugging tools are used with Visual Studio to optimize the code performance and get a better idea of what is happening ‘under the hood’ of the GPUs in real time.
NVIDIA: What approaches did the software team find useful for CUDA development?
Adam: Much of the initial focus has been on translating existing MATLAB scripts to CUDA C++. Our engineers first wrote the sequential C++ translation of the MATLAB script, and from there wrote the equivalent CUDA C++ code. That way the results from all three implementations could be compared for accuracy and running time.
In general, we found that the CUDA convex optimization implementations scale very well, and run significantly faster than both sequential C++ and MATLAB. The fact that there is a fast interface between MATLAB and CUDA means that we have a number of viable options when it comes to development, and can selectively export the heavy lifting to the GPU. [Ed: for more on GPU computing in MATLAB, read our post “Prototyping Algorithms and Testing CUDA Kernels in MATLAB“.]
NVIDIA: What types of parallel algorithms are being implemented?
Adam: For the signal processing and machine learning aspects of our research, we make extensive use of BLAS and LAPACK libraries for solving systems of equations and performing other linear algebraic operations. We also routinely use FFTs for spectral analysis.
CUDA has proved invaluable in accelerating these processes. In particular, we have developed a CUDA implementation of the Alternating Direction Method of Multipliers (ADMM) algorithm (Boyd et al, 2011) for distributed convex optimization. Solutions to many biophysical inverse problems we deal with (source localization, dynamical system identification, etc.) can be approximated by use of regularized and/or sparse linear regression. ADMM is well-suited for such problems, and with CUDA we can substantially scale up the size/complexity of the models while maintaining performance suitable for real-time analysis.
In addition, we are finishing work on a CUDA implementation of a second-order cone program (SOCP), an additional convex optimization which we will be using to implement a fast l1l2-norm inverse solver.
Also, our engineering team has written hybrid CPU/GPU implementations of algorithms useful for directed acyclic graphs. Since many brain-related data sets can be modeled as graphs, there is much new work that can and will be done in this area.
The libraries used were cuBLAS, cuSPARSE and Thrust. For the sensor-based EEG work, the matrices tend to be dense so cuBLAS was preferred. The cuSPARSE library was used to convert some of the native MATLAB sparse formats into the cuBLAS dense formats, because that was the fastest method of format conversion available. We did have to write a number of custom CUDA C++ algorithms for our work, like a dense Cholesky factoring kernel.
NVIDIA: If you had more computing power, what could you do?
Adam: Many of the present challenges in real-time, non-invasive brain monitoring amount to accurately inferring (i.e. learning, or identifying) the parameters of statistical models of ongoing brain activity from very limited amounts of observed data.
To achieve this goal, we must optimize very large numbers of parameters, which can be computationally prohibitive on ordinary serial computing architectures. With increased computing power afforded by parallel computing architectures, we can dramatically scale up the complexity of our models—both in terms of the number of model parameters as well as the computational complexity of the algorithms used to find an optimal solution—without sacrificing real-time processing capabilities.
Much of the real-time signal processing and machine learning we are developing for EEG can be extended to modeling and visualizing brain dynamics from this much richer source of information. Of course, with the increased data dimensionality, the complexity of our models will likewise increase (consider the challenge of learning the parameters of a neural network with 1 million “neurons” versus a network with only 10 neurons).
Powerful computing capabilities will be essential for making such inferences tractable, particularly for real-time monitoring. A further consideration is the use of “Big Data” — electrophysiological and behavioral data collected from a large number (hundreds, or thousands) of individuals — to obtain informative priors and constraints that can improve modeling of a single individual’s brain dynamics and activity, as well as improve decoding of one’s mental states, detection of clinical pathology and brain disease, and more. Analyzing and modeling such large amounts of data also poses a major computational challenge, which can be addressed by more powerful computing capabilities.
NVIDIA: What are the biggest challenges going forward?
Adam: Well, the present challenges are many. Some of the most critical involve development and application of novel statistical learning approaches to more accurately identify high-resolution cortical source activity and dynamical interactions from limited amounts of EEG data recorded in noisy “real-world” environments.
However, these algorithms must be capable of operating efficiently, producing accurate inferences of complex models with minimal processing delay. This is a computing challenge which can be addressed by advances both in distributed optimization algorithms as well as in parallel computing architectures.
The performance of the Kepler generation of GPUs has been impressive, and fortunately there is a helpful community of CUDA developers who actively share source code and advice. Nothing worthwhile is easy, but challenges are better overcome by the collaborative open-source software development model.
NVIDIA: What are you most excited about, in terms of advancements over the next decade?
Adam: The human brain is a highly dynamic system, constantly changing, adapting to meet the environmental challenges of the moment. The brain is also a distributed information processing system, wherein information is continually routed between multiple brain regions, acting together to process information.
To truly understand the brain — both in health and in disease — we have to be able to track changes in such distributed activity on the time scale at which they are occurring. Furthermore, the ability to measure and model these dynamics in “real world” settings, outside artificially constrained laboratory environments, is critical to improving our understanding of natural human brain function and its relationship to cognition and behavior.
Further still, the ability to make such inferences in “real-time” is of immense value for practical applications of brain monitoring, including medical diagnostics and intervention, human-machine interfaces and neural prosthetics, and even social interaction and entertainment.
In our lab, we are now attempting to integrate real-time neural EEG data with adaptive video game mechanics, neurofeedback algorithms and transcranial electrical stimulation to create a closed-loop system to accelerate the time-course of learning and repair.
The hardware and software tools necessary to achieve these goals are rapidly advancing, and improvements in computing capabilities is a key force in driving this advancement. Taken together, advances over the next decade in high-resolution, real-time brain monitoring, will have a profound positive impact on future technologies in multiple domains, from redefining medicine and personalized health care to transforming the way in which humans interact with each other and with machines in our lives.
NVIDIA has done a great job of demonstrating this new technology, and providing support to researchers using CUDA. Given that there are many talented engineers and researchers using this technology, it seems probable that this will lead to significant advancements in a number of scientific fields. It could be argued that we are entering into the ‘Golden Age’ of GPU computing, and I believe that our lab will be able to improve our understanding of brain activity through this technology.
Read more GPU Computing Spotlights.