CUDA
Apr 19, 2024
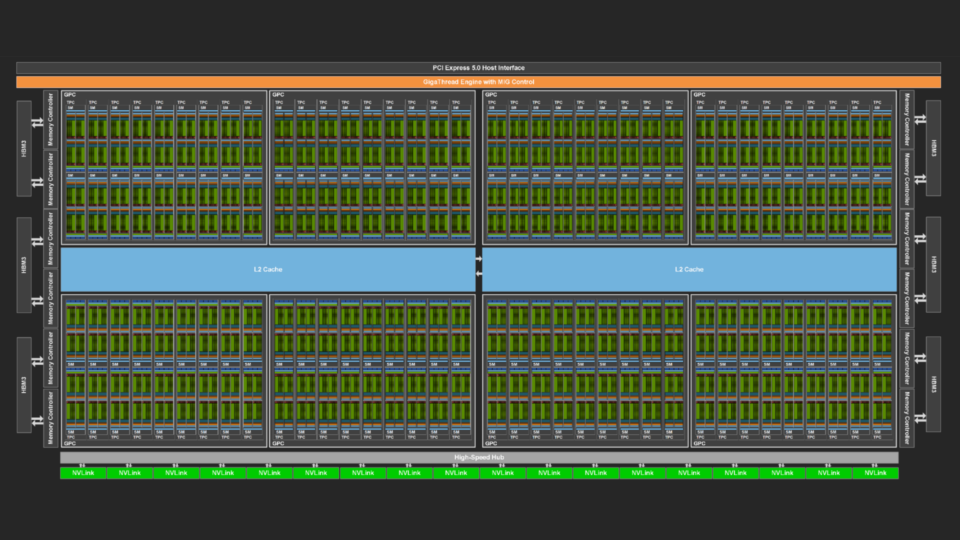
Measuring the GPU Occupancy of Multi-stream Workloads
NVIDIA GPUs are becoming increasingly powerful with each new generation. This increase generally comes in two forms. Each streaming multi-processor (SM), the...
11 MIN READ

Mar 27, 2024
Efficient CUDA Debugging: Using NVIDIA Compute Sanitizer with NVIDIA Tools Extension and Creating Custom Tools
NVIDIA Compute Sanitizer is a powerful tool that can save you time and effort while improving the reliability and performance of your CUDA applications....
14 MIN READ

Mar 25, 2024
Building High-Performance Applications in the Era of Accelerated Computing
AI is augmenting high-performance computing (HPC) with novel approaches to data processing, simulation, and modeling. Because of the computational requirements...
6 MIN READ
Mar 14, 2024
Just Released: NVIDIA cuSPARSELt 0.6
NVIDIA cuSPARSELt harnesses Sparse Tensor Cores to accelerate general matrix multiplications. Version 0.6. adds support for the NVIDIA Hopper architecture.
1 MIN READ
Mar 06, 2024
CUDA Toolkit 12.4 Enhances Support for NVIDIA Grace Hopper and Confidential Computing
The latest release of CUDA Toolkit, version 12.4, continues to push accelerated computing performance using the latest NVIDIA GPUs. This post explains the new...
9 MIN READ

Feb 28, 2024
Optimizing OpenFold Training for Drug Discovery
Predicting 3D protein structures from amino acid sequences has been an important long-standing question in bioinformatics. In recent years, deep...
7 MIN READ
Jan 12, 2024
Just Released: cuBLASDx
cuBLASDx allows you to perform BLAS calculations inside your CUDA kernel, improving the performance of your application. Available to download in Preview...
1 MIN READ
Jan 05, 2024
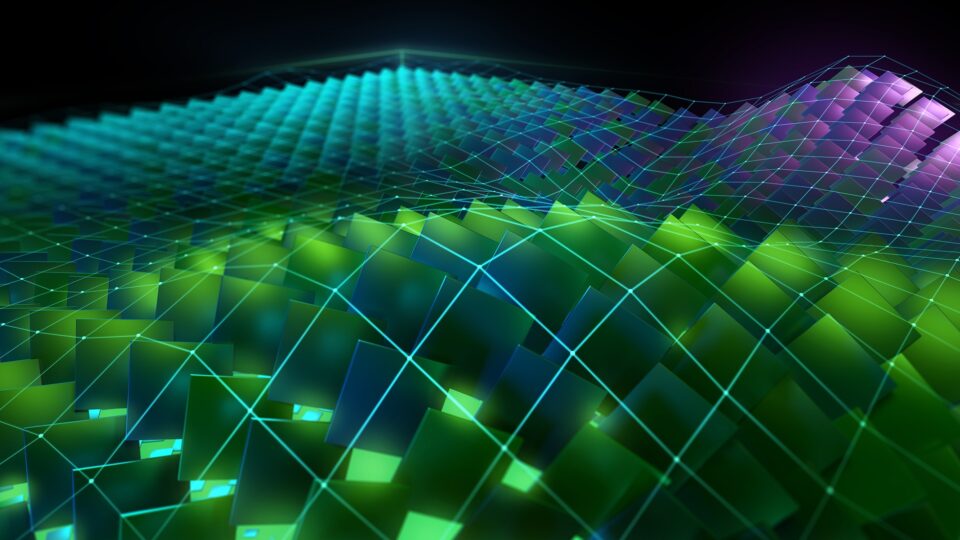
Improving CUDA Initialization Times Using cgroups in Certain Scenarios
Many CUDA applications running on multi-GPU platforms usually use a single GPU for their compute needs. In such scenarios, a performance penalty is paid by...
5 MIN READ
Dec 20, 2023
Just Released: cuBLASMp
cuBLASMp is a high-performance, multi-process, GPU-accelerated library for distributed basic dense linear algebra. It is available to download in Preview now.
1 MIN READ

Nov 29, 2023
CUDA-Q 0.5 Delivers New Features for Quantum-Classical Computing
NVIDIA CUDA-Q is a platform for building quantum-classical computing applications. It is an open-source programming model for heterogeneous computing such as...
4 MIN READ

Nov 21, 2023
Unlocking GPU Intrinsics in HLSL
There are some useful intrinsic functions in the NVIDIA GPU instruction set that are not included in standard graphics APIs. Updated from the original 2016 post...
9 MIN READ

Nov 17, 2023
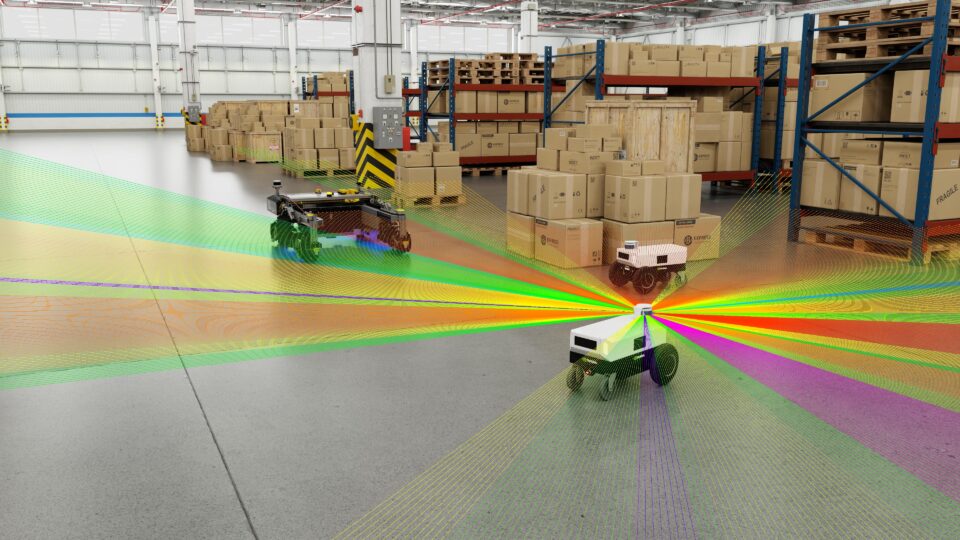
Boosting Custom ROS Graphs Using NVIDIA Isaac Transport for ROS
NVIDIA Isaac Transport for ROS (NITROS) is the implementation of two hardware-acceleration features introduced with ROS 2 Humble-type adaptation and type...
11 MIN READ

Nov 16, 2023
Unlock the Power of NVIDIA Grace and NVIDIA Hopper Architectures with Foundational HPC Software
High-performance computing (HPC) powers applications in simulation and modeling, healthcare and life sciences, industry and engineering, and more. In the modern...
7 MIN READ

Nov 07, 2023
CUDA-Accelerated Robot Motion Generation in Milliseconds with NVIDIA cuRobo
Real-time autonomous robot navigation powered by a fast motion-generation algorithm can enable applications in several industries such as food and services,...
3 MIN READ

Nov 06, 2023
ICYMI: Leveraging the Power of GPUs with CuPy in Python
See how KDNuggets achieved 500x speedup using CuPy and NVIDIA CUDA on 3D arrays.
1 MIN READ
Nov 01, 2023
CUDA Toolkit 12.3 Delivers New Features for Accelerated Computing
The latest release of CUDA Toolkit continues to push the envelope of accelerated computing performance using the latest NVIDIA GPUs. New features of this...
4 MIN READ