Thrust
Mar 18, 2015
The Power of C++11 in CUDA 7
Today I'm excited to announce the official release of CUDA 7, the latest release of the popular CUDA Toolkit. Download the CUDA Toolkit version 7 now from CUDA...
12 MIN READ

Feb 12, 2015
Accelerating Bioinformatics with NVBIO
NVBIO is an open-source C++ template library of high performance parallel algorithms and containers designed by NVIDIA to accelerate sequence analysis and...
11 MIN READ

Jan 13, 2015
CUDA 7 Release Candidate Feature Overview: C++11, New Libraries, and More
It's almost time for the next major release of the CUDA Toolkit, so I'm excited to tell you about the CUDA 7 Release Candidate, now available to all CUDA...
9 MIN READ

Apr 10, 2014
CUDA Spotlight: GPU-Accelerated Agent-Based Simulation of Complex Systems
This week’s Spotlight is on Dr. Paul Richmond, a Vice Chancellor's Research Fellow at the University of Sheffield (a CUDA Research Center). Paul's research...
5 MIN READ

Feb 06, 2014
CUDACasts Episode 16: Thrust Algorithms and Custom Operators
Continuing the Thrust mini-series (see Part 1), today's episode of CUDACasts focuses on a few of the algorithms that make Thrust a flexible and powerful...
1 MIN READ

Jan 29, 2014
CUDACasts Episode 15: Introduction to Thrust
Whenever I hear about a developer interested in accelerating his or her C++ application on a GPU, I make sure to tell them about Thrust. Thrust is a parallel...
1 MIN READ

Oct 08, 2013
CUDA Spotlight: GPU-Accelerated Genomics
This week's Spotlight is on Dr. Knut Reinert. Knut is a professor at Freie Universität in Berlin, Germany, and chair of the Algorithms in Bioinformatics group...
7 MIN READ

Apr 18, 2013
Copperhead: Data Parallel Python
Programming environments like C and Fortran allow complete and unrestricted access to computing hardware, but often require programmers to understand the...
12 MIN READ
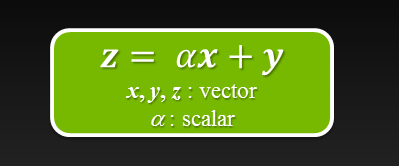
Jul 02, 2012
Six Ways to SAXPY
For even more ways to SAXPY using the latest NVIDIA HPC SDK with standard language parallelism, see N Ways to SAXPY: Demonstrating the Breadth of GPU...
8 MIN READ

Jun 06, 2012
Expressive Algorithmic Programming with Thrust
Thrust is a parallel algorithms library which resembles the C++ Standard Template Library (STL). Thrust's High-Level interface greatly enhances...
9 MIN READ