Spotlight

Mar 05, 2024
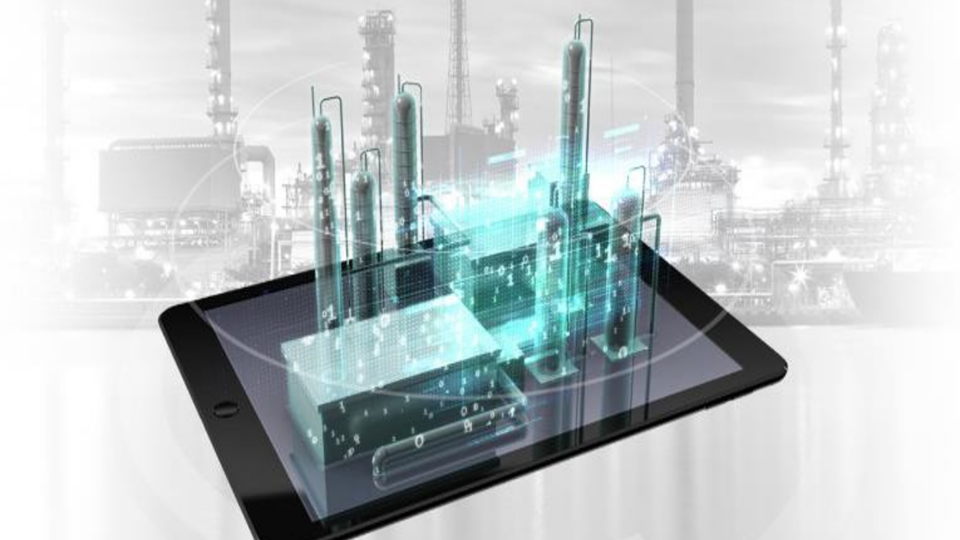
Spotlight: Honeywell Accelerates Industrial Process Simulation with NVIDIA cuDSS
For over a decade, traditional industrial process modeling and simulation approaches have struggled to fully leverage multicore CPUs or acceleration devices to...
8 MIN READ

Feb 28, 2024
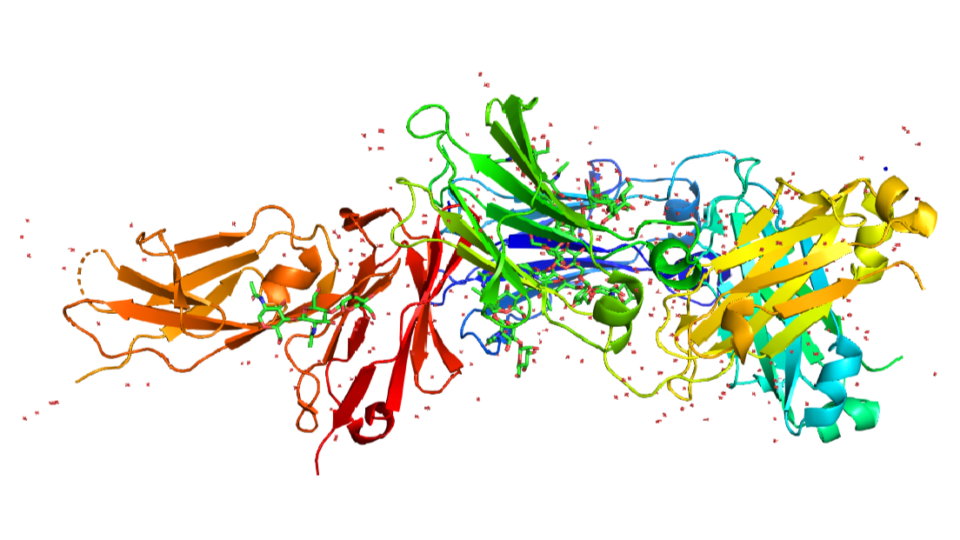
Optimizing OpenFold Training for Drug Discovery
Predicting 3D protein structures from amino acid sequences has been an important long-standing question in bioinformatics. In recent years, deep...
7 MIN READ

Feb 22, 2024
Enhance Immersive Experiences with the New Varjo XR-4 Series Headsets, Powered by NVIDIA
Developers and enterprises can now deploy lifelike virtual and mixed reality experiences with Varjo's latest XR-4 series headsets, which are integrated with...
3 MIN READ

Feb 21, 2024
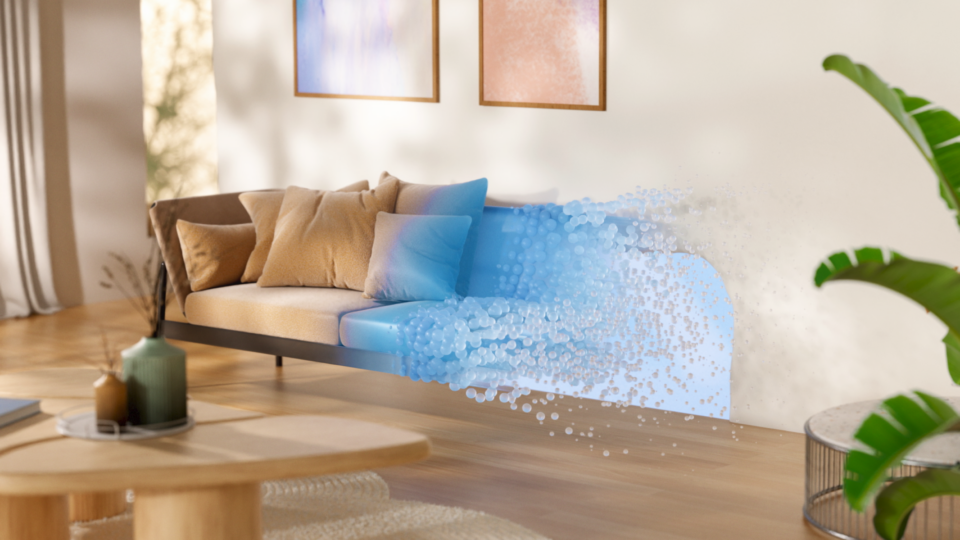
Spotlight: HOMEE AI Delivers AI-Powered Spatial Planning to Your Living Room
HOMEE AI, an NVIDIA Inception member based in Taiwan, has developed an “AI-as-a-service” spatial planning solution to disrupt the $650B global home decor...
7 MIN READ

Jan 09, 2024
Enhancing Phone Customer Service with ASR Customization
At the core of understanding people correctly and having natural conversations is automatic speech recognition (ASR). To make customer-led voice assistants and...
7 MIN READ

Dec 19, 2023
Breakthrough in Functional Annotation with HiFi-NN
Enzymes are vital biological catalysts for a multitude of processes, from cellular metabolism to industrial manufacturing. The applications of artificial...
7 MIN READ
Nov 29, 2023
Boost Meeting Productivity with AI-Powered Note-Taking and Summarization
Meetings are the lifeblood of an organization. They foster collaboration and informed decision-making. They eliminate silos through brainstorming and...
6 MIN READ
Nov 09, 2023
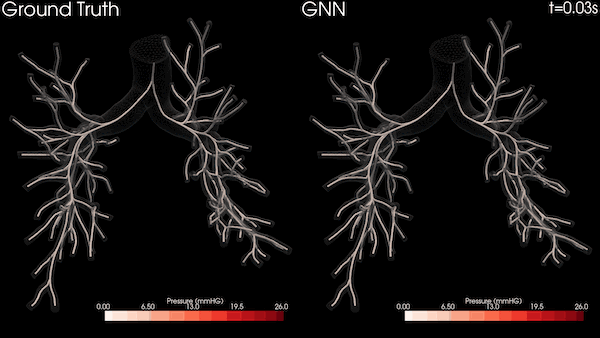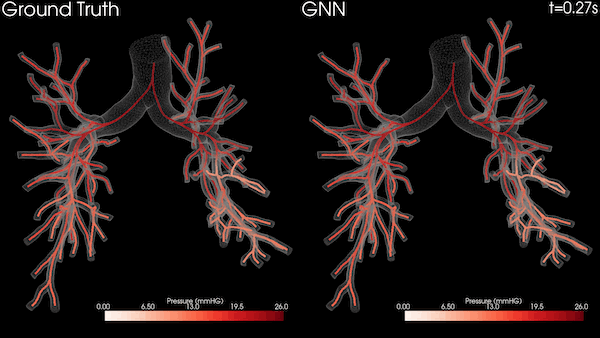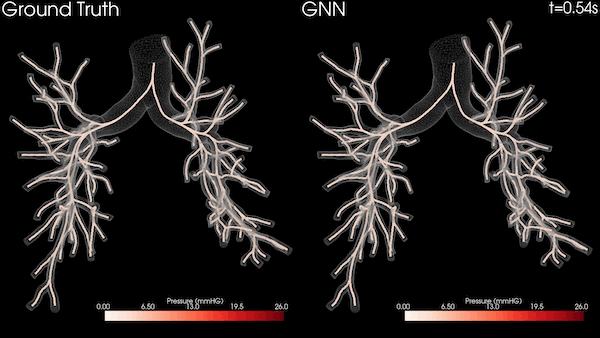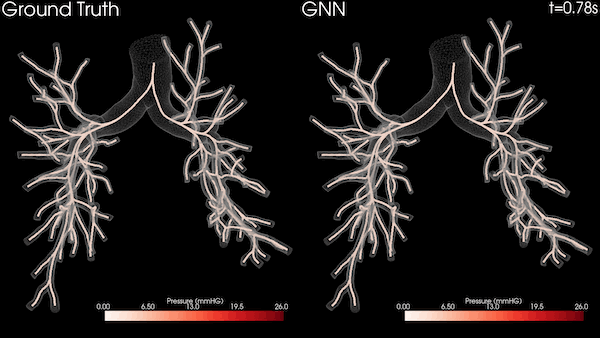
Enabling Greater Patient-Specific Cardiovascular Care with AI Surrogates
A Stanford University team is transforming heart healthcare with near real-time cardiovascular simulations driven by the power of AI. Harnessing...
8 MIN READ

Nov 09, 2023
Accelerating Neurosymbolic AI with RAPIDS and Prometheux Vadalog Parallel
As the scale of available data continues to grow, so does the need for scalable and intelligent data processing systems to swiftly harness useful knowledge....
11 MIN READ

Nov 08, 2023
Whole Human Brain Neuro-Mapping at Cellular Resolution on NVIDIA DGX
Whole human brain imaging of 100 brains at a cellular level within a 2-year timespan, and subsequent analysis and mapping, requires accelerated supercomputing...
9 MIN READ

Nov 03, 2023
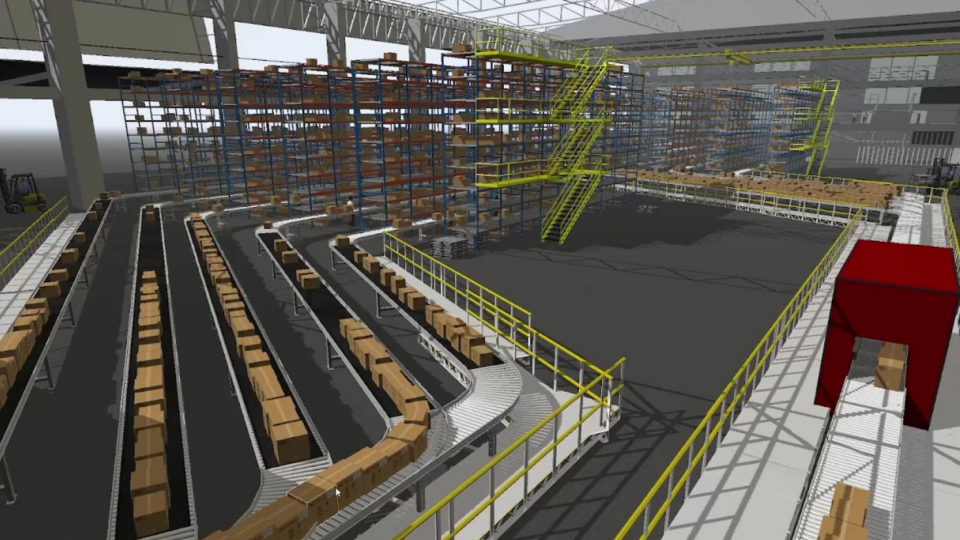
Analyze, Visualize, and Optimize Real-World Processes with OpenUSD in FlexSim
For manufacturing and industrial enterprises, efficiency and precision are essential. To streamline operations, reduce costs, and enhance productivity,...
6 MIN READ

Oct 28, 2023
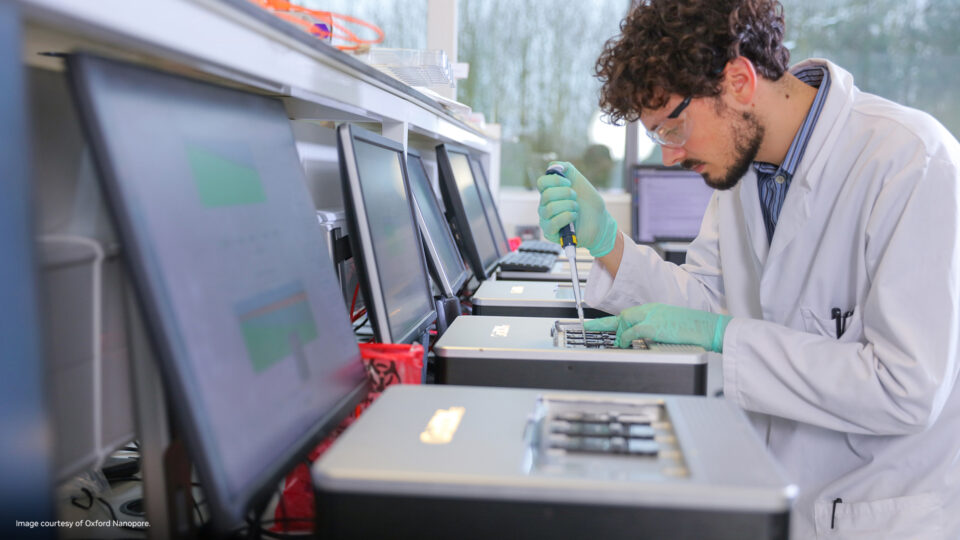
Accelerate Genomic Analysis for Any Sequencer with NVIDIA Parabricks v4.2
Parabricks version 4.2 has been released, furthering its mission to deliver unprecedented speed, cost-effectiveness, and accuracy in genomics sequencing...
7 MIN READ

Oct 13, 2023
Supercharge Graph Analytics at Scale with GPU-CPU Fusion for 100x Performance
Graphs form the foundation of many modern data and analytics capabilities to find relationships between people, places, things, events, and locations across...
11 MIN READ
Oct 02, 2023
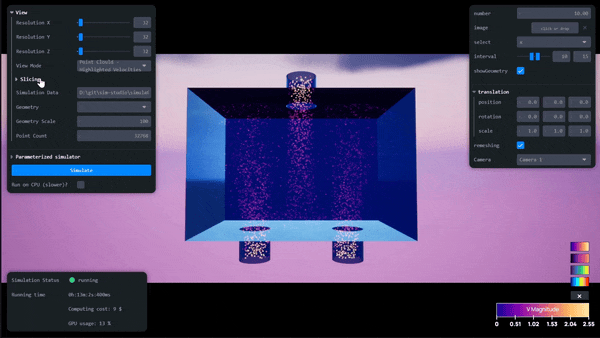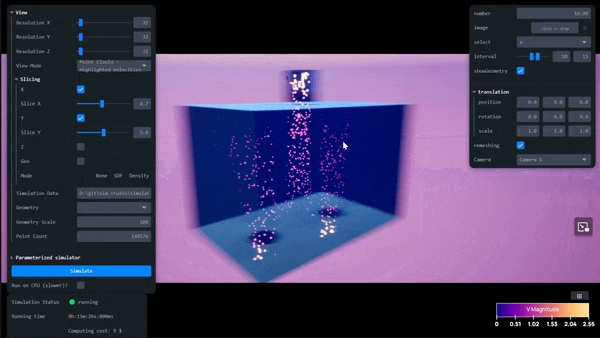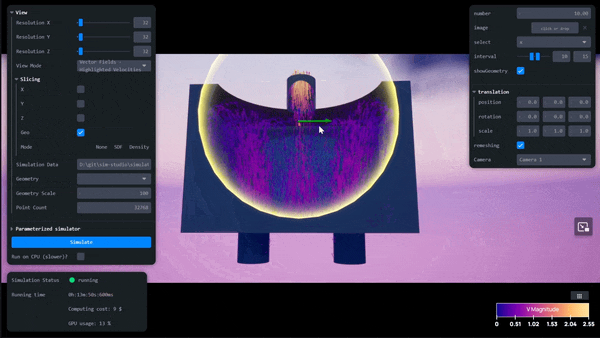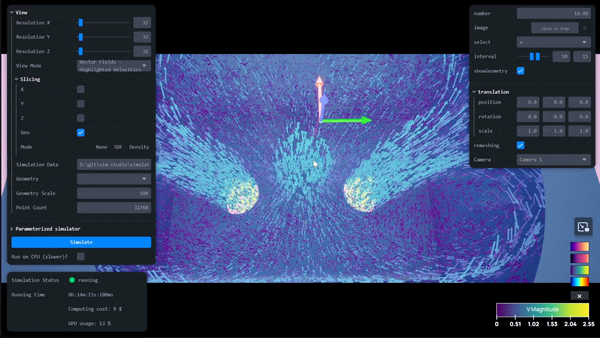
AI-Powered Simulation Tools for Surrogate Modeling Engineering Workflows with Siml.ai and NVIDIA Modulus
Simulations are quintessential for complex engineering challenges, like designing nuclear fusion reactors, optimizing wind farms, developing carbon capture and...
6 MIN READ

Sep 11, 2023
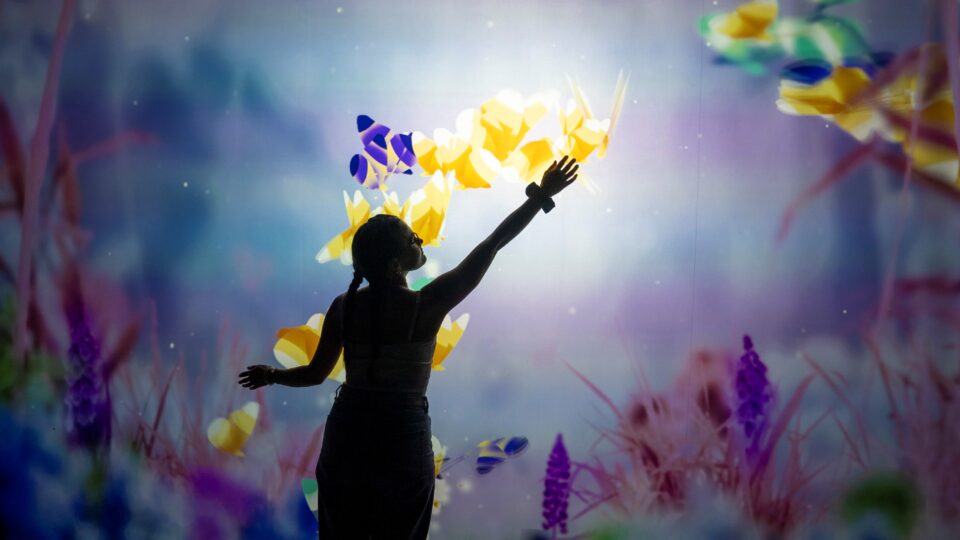
Creating Immersive Events with OpenUSD and Digital Twins
Moment Factory is a global multimedia entertainment studio that combines specializations in video, lighting, architecture, sound, software, and interactivity to...
8 MIN READ

Aug 10, 2023
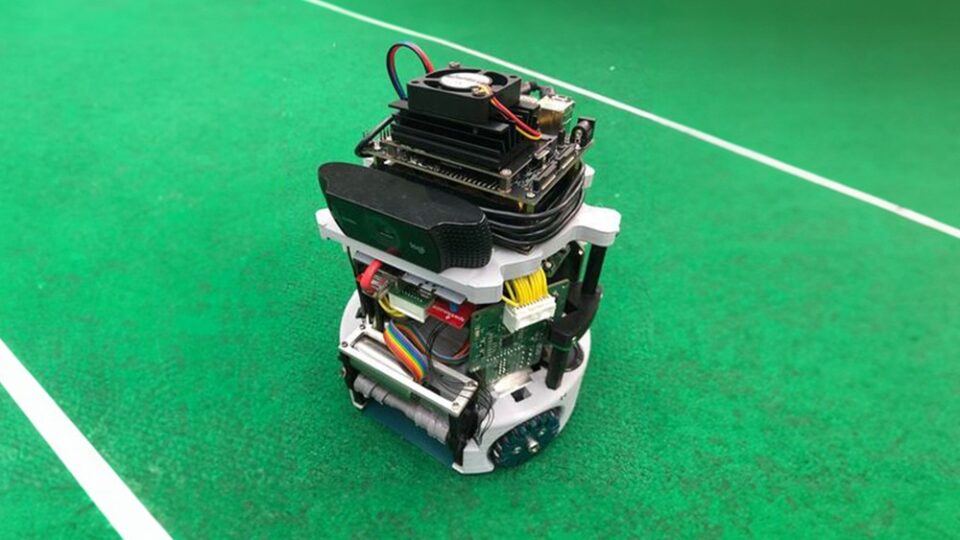
NVIDIA Jetson Project of the Month: This Autonomous Soccer Robot Can Aim, Shoot, and Score
Soccer is considered one of the most popular sports around the world. And with good reason: the action is often intense, and the game combines both physicality...
9 MIN READ