Python

Feb 21, 2024
NVIDIA TensorRT-LLM Revs Up Inference for Google Gemma
NVIDIA is collaborating as a launch partner with Google in delivering Gemma, a newly optimized family of open models built from the same research and technology...
4 MIN READ
Jan 30, 2024
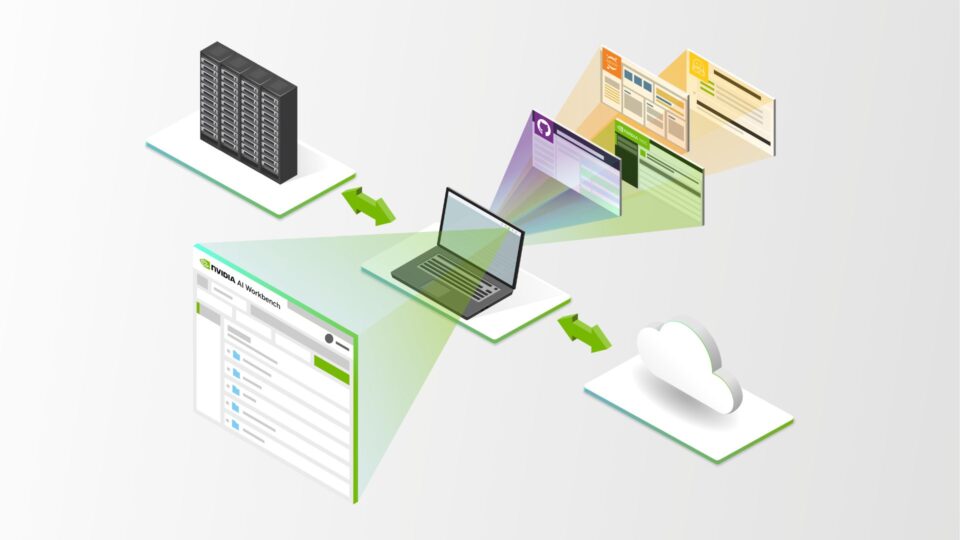
Create, Share, and Scale Enterprise AI Workflows with NVIDIA AI Workbench, Now in Beta
NVIDIA AI Workbench is now in beta, bringing a wealth of new features to streamline how enterprise developers create, use, and share AI and machine learning...
10 MIN READ

Nov 13, 2023
NVIDIA Deep Learning Institute Launches Science and Engineering Teaching Kit
AI is quickly becoming an integral part of diverse industries, from transportation and healthcare to manufacturing and finance. AI powers chatbots, recommender...
5 MIN READ

Nov 08, 2023
New Workshop: Rapid Application Development Using Large Language Models
Interested in developing LLM-based applications? Get started with this exploration of the open-source ecosystem.
1 MIN READ

Nov 06, 2023
ICYMI: Leveraging the Power of GPUs with CuPy in Python
See how KDNuggets achieved 500x speedup using CuPy and NVIDIA CUDA on 3D arrays.
1 MIN READ

Oct 11, 2023
Announcing NVIDIA SteerLM: A Simple and Practical Technique to Customize LLMs During Inference
With the advent of large language models (LLMs) such as GPT-3, Megatron-Turing, Chinchilla, PaLM-2, Falcon, and Llama 2, remarkable progress in natural language...
10 MIN READ

Sep 21, 2023
Just Released: NVIDIA Modulus 23.09
NVIDIA Modulus 23.09 is now available, providing ease-of-use updates, fixes, and other enhancements.
1 MIN READ

Aug 30, 2023
New Video Tutorial: Profiling and Debugging NVIDIA CUDA Applications
Episode 5 of the NVIDIA CUDA Tutorials Video series is out. Jackson Marusarz, product manager for Compute Developer Tools at NVIDIA, introduces a suite of tools...
2 MIN READ
Aug 22, 2023
Simplifying GPU Application Development with Heterogeneous Memory Management
Heterogeneous Memory Management (HMM) is a CUDA memory management feature that extends the simplicity and productivity of the CUDA Unified Memory programming...
16 MIN READ
Aug 08, 2023
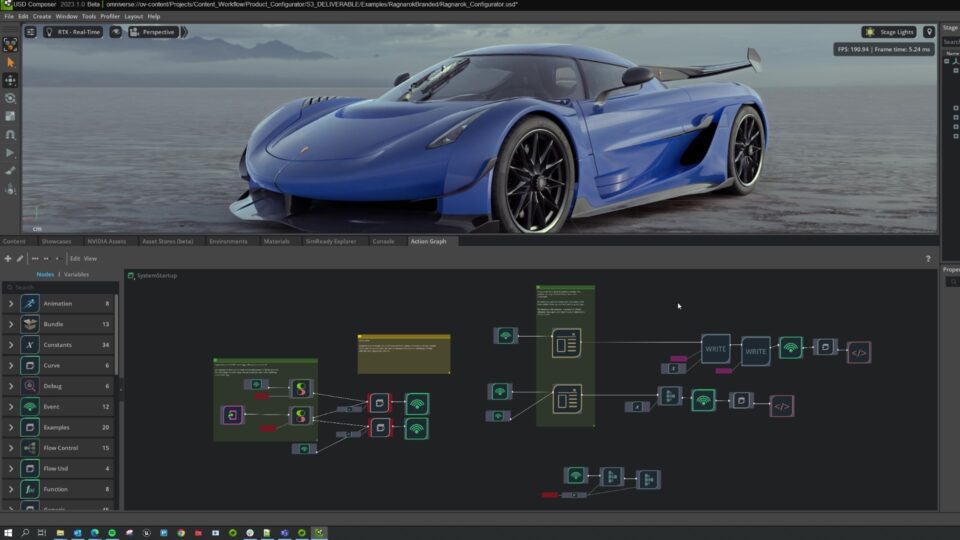
Accelerate 3D Workflows with Modular, OpenUSD-Powered Omniverse Release
The latest release of NVIDIA Omniverse delivers an exciting collection of new features based on Omniverse Kit 105, making it easier than ever for developers to...
7 MIN READ
Aug 03, 2023
Securing LLM Systems Against Prompt Injection
Prompt injection is a new attack technique specific to large language models (LLMs) that enables attackers to manipulate the output of the LLM. This attack is...
15 MIN READ
Jun 28, 2023
How to Deploy an AI Model in Python with PyTriton
AI models are everywhere, in the form of chatbots, classification and summarization tools, image models for segmentation and detection, recommendation models,...
6 MIN READ
Jun 27, 2023
GPU-Accelerated Single-Cell RNA Analysis with RAPIDS-singlecell
Single-cell sequencing has become one of the most prominent technologies used in biomedical research. Its ability to decipher changes in the transcriptome and...
13 MIN READ
May 05, 2023
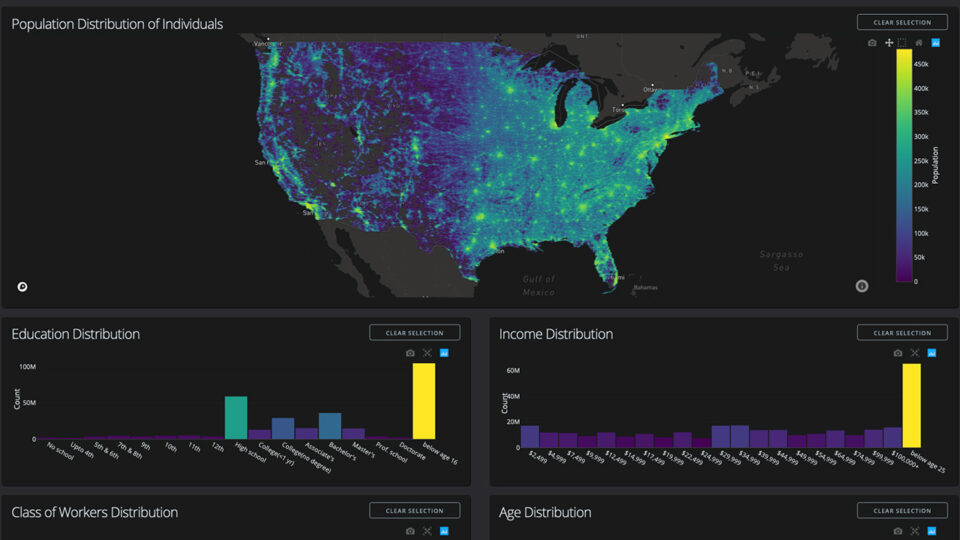
NVIDIA On-Demand: Top Data Science Sessions from GTC 2023
Learn from experts about how to optimize a data pipeline or use machine learning for anomaly detection with these 15 educational sessions.
1 MIN READ

Apr 26, 2023
A Comprehensive Guide to Interaction Terms in Linear Regression
Linear regression is a powerful statistical tool used to model the relationship between a dependent variable and one or more independent variables (features)....
13 MIN READ

Apr 25, 2023
Increasing Inference Acceleration of KoGPT with NVIDIA FasterTransformer
Transformers are one of the most influential AI model architectures today and are shaping the direction of future AI R&D. First invented as a tool for...
6 MIN READ